Researchers from DTU and Copenhagen University are the first to show that protein analysis at the single cell level in a tumor sample from an acute myeloid leukemia patient can be used to detect cancer stem cells that evade chemotherapy. The technology the researchers have developed can also be applied on biotechnological production where it can provide novel insights at the single-cell level, which in turn can be used to improve cell line production capacity.
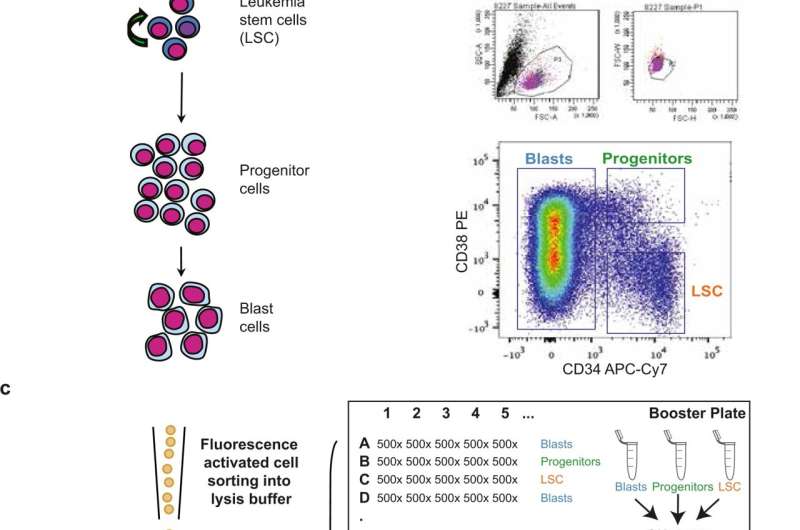
AML (acute myeloid leukemia) is a cancer of the blood that affects the normal stem cells in the bone marrow, which are thereby transformed into cancer stem cells, and maturation of the cells stops prematurely. These immature cells accumulate rapidly in the bone marrow and displace the normal cells. This leads to severe deficiency of normally functioning cells in the blood. AML is a very aggressive form of cancer and its treatment consists of intensive chemotherapy, which in many cases can minimize the amount of immature, diseased cells in the bone marrow to less than 5 percent. This is an indication that the disease is at rest and can no longer be detected by microscopic examination of the bone marrow and the patient is considered cured.
However, even though the disease cannot be clinically detected, there could still be cancer stem cells present in the bone marrow. With approximately 50% patient relapse and only 22% survival rate after five years of disease onset, this a real threat to a full recovery.
Analyzing cancer at the single-cell level
Researchers at DTU and Copenhagen University therefore decided to go a level deeper and examine cancer samples at the single cell level; through a series of optimizations, their workflow allows studying the proteomes of single cells without any necessary pre-enrichment. The proteome constitutes all the proteins in a cell, and it is the cellular proteins that are the cell’s workhorses. Thus, cellular proteomes represent high-resolution molecular maps of current cell states and are highly informative about cellular function. Through profiling of these protein signatures, it is now possible to distinguish cell types based on protein-level data only, and reveal the generally quiescent cancer stem cells.
“We want to target these quiescent leukemia stem cells, but the problem is that they constitute a very small part—less than 1% of the tumor samples. The standard approach of studying cancer, where mass spectrometry analysis is done on bulk tumor samples, misses these very minute cell populations. Now, for the first time, we have the chance to understand the proteins and how the protein signaling networks go wrong in those cancer cells that evade therapy,” says Associate Professor and Head of the Proteomics Core Erwin Schoof from DTU Bioengineering, who headed the study.
The study shows that single-cell proteomics is ready to be used to answer similar questions as current transcriptome-based (i.e. RNA) methods but with the added advantage of providing meaningful information about the actual protein expression within the otherwise hidden cancer cells. Something current bulk-level technologies cannot.
The underlying technology also allows the researchers to understand other cell systems in for example, various cancers and it can be used to provide novel insights in biotechnological production systems. For example, the technology can reveal which cell populations are good producers versus those that are not, and identify cellular markers to help select those high producers.
The challenge of the single-cell data is that it is very large and multidimensional. Normally, there would be two conditions to an analysis: cancer and no cancer and it could be a case of comparing a group of cancer samples to a group of non-cancer.
With the single-cell approach, there will now be tens of thousands of cells for each clinical sample, that will each have a long list of proteins and their expression levels. The complexity is so large that the researchers had to invent a new piece of software (SCeptre) that could help analyze and visualize the data for them. Erwin Schoof continues:
Source: Read Full Article
